
I am an assistant professor of chemistry at the Technical University of Denmark (DTU). I am a member of the European Laboratory for Learning and Intelligent Systems (ELLIS), a member of the Pioneer Centre for AI in Copenhagen, and a member of the Center for Translational Protein Design (CTPD) at DTU. I also serve part-time as Director of Machine Learning Research at Jura Bio.
I work on fundamental machine learning methodology for molecules. I develop probabilistic and generative methods to steer chemical synthesis and high-throughput measurements, and to learn from natural experiments in humans and the environment. I aim to advance our basic understanding of how to learn about the molecular world, from both an applied and a theoretical perspective.
Previously, I was a postdoctoral research scientist with David Blei in the Data Science Institute at Columbia University. I received my PhD in Biophysics from Harvard University in 2022, advised by Debora Marks and Jeff Miller, as a Hertz Foundation Fellow. I received my A.B. in Chemistry and Physics with highest honors from Harvard in 2016, working with Adam Cohen.
Please reach out if you are interested in working with me or collaborating.
Machine Learning for Molecules
Our research is on fundamental machine learning methodology for molecules. We develop probabilistic and generative methods to steer chemical synthesis and high-throughput screening, and to learn from natural experiments in humans and the environment.
We aim to advance our basic understanding of how to learn about the molecular world. How can we estimate molecules' effects on complex biological systems? How can we design and synthesize molecules with desired properties? How can we traverse chemical space to find useful molecules?
We investigate these questions from both an applied and a theoretical perspective.
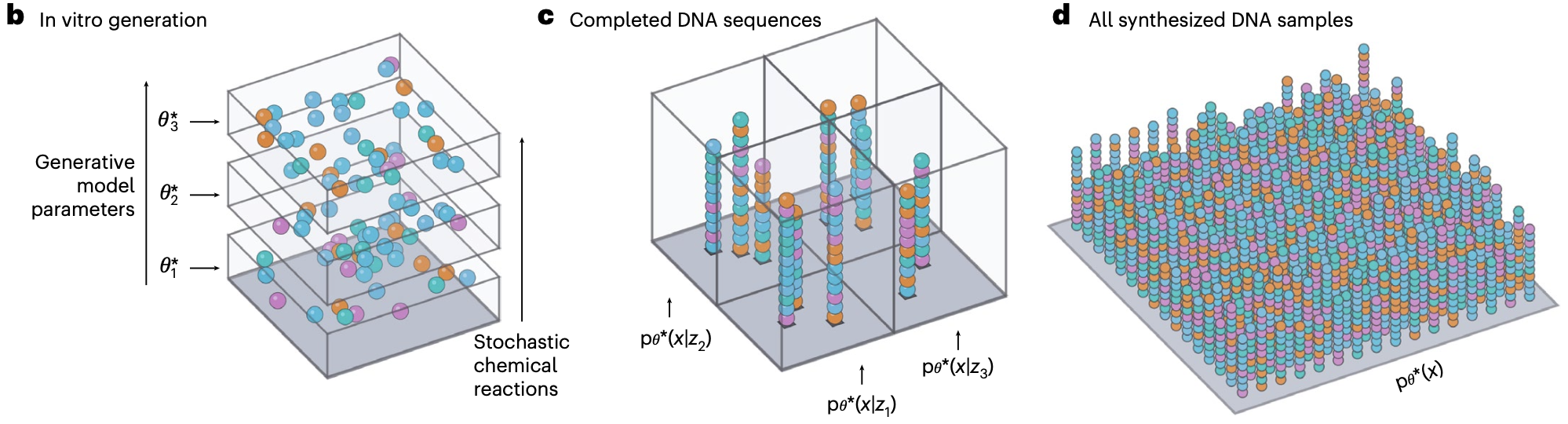
One major line of research is on the co-design of experiments and inference algorithms. We develop new ML methods to actively steer chemical synthesis, optimize high-throughput testing, and improve model learning. This tight integration of laboratory data generation and algorithmic development accelerates molecular discovery. We apply these techniques to design and discover therapeutics, enzymes and more.
Our work intersects with experimental design, generative modeling, Bayesian optimization, and laboratory automation. We are especially motivated by a desire to understand how the unique properties of molecular systems might let us overcome the limits of conventional ML experimental design and optimization. We develop algorithms that can exploit randomness, stochasticity and heterogeneity at the molecular level to pack more information into experiments. We use these techniques to overcome the unique learning challenges of molecular discovery, including rare properties, sparse structure-activity relationships, and discrete combinatorial spaces.
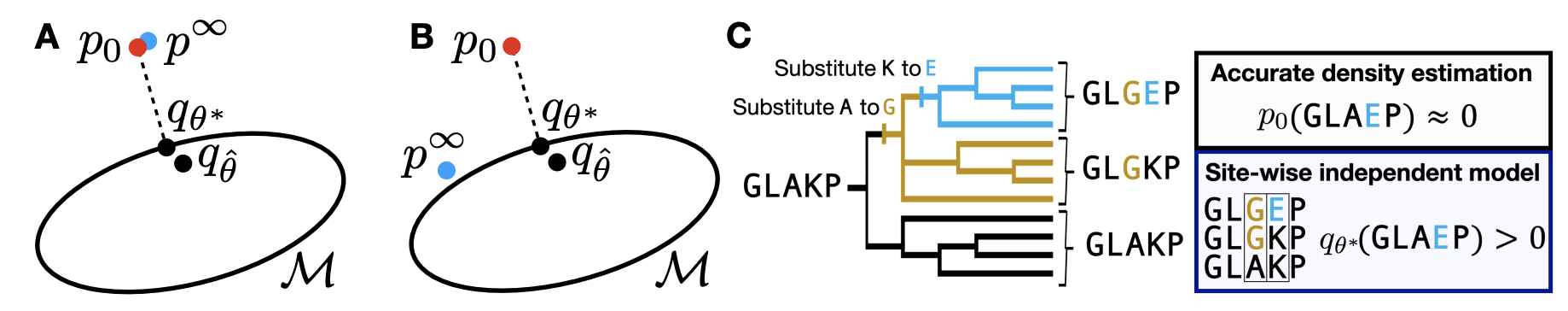
A second major line of research learns from natural experiments, outside the controlled environment of the laboratory. This includes large-scale evolutionary processes, environmental samples and patient-derived data. Molecules made in nature often have properties that are difficult to reproduce synthetically, such as the bioactivity of natural products or the function of proteins in the human body. We develop ML methods that can learn from natural experiments to design molecules with desired properties.
Our work intersects with generative models, causal inference, and human and evolutionary biology. Generative models have proven to be powerful tools for learning the constraints on molecules imposed by evolution, but their inferences are biased, imperfect and incomplete. We investigate methods that can exploit the scale and diversity of natural experiments to learn the true range of molecular possibilities, and to align flexible generative models with underlying chemistry and biology. We aim to use natural experiments to systematically bridge generalization gaps between the laboratory and the outside world, such as the valley of death between preclinical experiments and clinical trials in humans.
*Equal contribution · Google Scholar

Affiliations: DTU Department of Chemistry · Jura Bio
Office: Building 206, Room 258, Kemitorvet, 2800 Kgs. Lyngby
Email: enawe [at] dtu.dk
Brief bio (plain text)

I am always looking for motivated students, postdocs, and collaborators interested in probabilistic machine learning and its applications to the molecular sciences. Please reach out if you are interested: enawe [at] dtu.dk

ML and Molecules Reading Group — Spring 2026
The ML & Molecules reading group covers fundamental machine learning methodology and its intersection with the molecular sciences. This semester we are covering two interrelated topics: first, the interface of machine learning and simulation, and second, experimental design and planning in the time domain. Our goal is to understand how to model the dynamics of chemical and biological systems, and their response to perturbation. Large scale computing has transformed scientific discovery both through physical simulation and through machine learning. Yet these two forms of scientific computing sit in uneasy relationship, operating with distinct logic and methodology. We will investigate the interface between machine learning and simulation. Next, we turn our attention to the implications of these models for experimental design. Current machine learning techniques for molecular discovery typically rely on separate controlled experiments of different molecules. These techniques are often bottlenecked in throughput. We study alternative experimental designs that test the effects of different interventions by turning them on or off over time. In these experiments, the time dynamics of the system set the speed of information gain. To analyze these experiments, we must understand systems' response to perturbations, and leverage them to create new experimental protocols.
Sign up for the mailing list
Noé et al. 2019. Boltzmann generators: Sampling equilibrium states of many-body systems with deep learning. link
Hyvärinen. 2005. Estimation of non-normalized statistical models by score matching. link
Plainer et al. 2025. Consistent Sampling and Simulation: Molecular Dynamics with Energy-Based Diffusion Models link
Mohamed and Lakshminarayanan. 2016. Learning in implicit generative models. link
Kabylda et al. 2025. Molecular Simulations with a Pretrained Neural Network and Universal Pairwise Force Fields. link
Papamakarios and Murray. 2016. Fast ε-free Inference of Simulation Models with Bayesian Conditional Density Estimation. link
Glynn et al. 2020. Adaptive Experimental Design with Temporal Interference: A Maximum Likelihood Approach. link
Liang and Recht. 2023. Randomization Inference When N Equals One. link
Schiebinger et al. 2019. Optimal-Transport Analysis of Single-Cell Gene Expression Identifies Developmental Trajectories in Reprogramming. link
Lorch et al. 2026. Latent Causal Diffusions for Single-Cell Perturbation Modeling. link
Lopez et al. 2023. Learning Causal Representations of Single Cells via Sparse Mechanism Shift Modeling. link
Raissi et al. 2017. Physics Informed Deep Learning (Part I): Data-driven Solutions of Nonlinear Partial Differential Equations. link
Contact: enawe [at] dtu.dk

ML and Molecules Reading Group — Fall 2025
Machine learning and AI have transformed scientific data analysis. But the experiments that generate data are usually taken as given. Better algorithmic control of scientific experiments could lead to better learning, enabling the next generation of scientific AI.
This semester, we studied the fundamental theory of experimental design. We read foundational statistical ideas about how to quantify the amount of information an experiment provides, studied emerging ML methods and algorithms for optimizing experimental designs, and looked at how these ideas are being applied in science, with a focus on protein design and engineering.
Sign up for the mailing list
Reading List
D.V. Lindley. 1956. On a Measure of the Information Provided by an Experiment. link
Romero et al. 2012. Navigating the protein fitness landscape with Gaussian processes. link
Frey et al. 2025. Lab-in-the-loop therapeutic antibody design with deep learning. link
Foster et al. 2020. A Unified Stochastic Gradient Approach to Designing Bayesian-Optimal Experiments. link
Krishnan et al. 2025. A generative deep learning approach to de novo antibiotic design. link
Smith et al. 2023. Prediction Oriented Bayesian Active Learning. link
Liu et al. 2024. Scalable, compressed phenotypic screening using pooled perturbations. link
Grover and Ermon. 2019. Uncertainty Autoencoders. link
Russo and van Roy. 2014. Learning to Optimize via Information-Directed Sampling. link
Huszar and Duvenaud. 2012. Optimally-Weighted Herding is Bayesian Quadrature. link
Contact: enawe [at] dtu.dk